附录 C — 贝叶斯计算
C.1 广义线性模型
先介绍泊松广义线性模型,包括模拟和计算,并和 Stan 实现的结果比较。
泊松广义线性模型如下:
\[ \begin{aligned} \log(\lambda) &= \beta_0 + \beta_1 x_1 + \beta_2 x_2 \\ Y &\sim \mathrm{Poisson}(u\lambda) \end{aligned} \]
设定参数向量 \(\beta = (\beta_0, \beta_1, \beta_2) = (0.5, 0.3, 0.2)\),观测变量 \(X_1\) 和 \(X_2\) 的均值都为 0,协方差矩阵 \(\Sigma\) 为
\[ \left[ \begin{matrix} 1.0 & 0.8 \\ 0.8 & 1.0 \end{matrix} \right] \]
C.2 生成模拟数据
模拟观测到的响应变量值和协变量值,添加漂移项
拟合泊松回归模型
fit_poisson_glm <- glm(y ~ X, family = poisson(link = "log"), offset = log(u))
summary(fit_poisson_glm)
Call:
glm(formula = y ~ X, family = poisson(link = "log"), offset = log(u))
Coefficients:
Estimate Std. Error z value Pr(>|z|)
(Intercept) 0.488932 0.009427 51.86 <2e-16 ***
X1 0.289984 0.014298 20.28 <2e-16 ***
X2 0.214846 0.014420 14.90 <2e-16 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
(Dispersion parameter for poisson family taken to be 1)
Null deviance: 6052.9 on 2499 degrees of freedom
Residual deviance: 2675.5 on 2497 degrees of freedom
AIC: 10773
Number of Fisher Scoring iterations: 4C.3 拟合泊松模型
下面准备数据
编译模型,抽样获取参数的后验分布
# 加载 cmdstanr 包
library(cmdstanr)
# 编译模型
mod_poisson <- cmdstan_model(
stan_file = "code/poisson_log_glm.stan",
compile = TRUE,
cpp_options = list(stan_threads = TRUE)
)
# 采样拟合模型
fit_poisson_stan <- mod_poisson$sample(
data = poisson_d, # 观测数据
init = inits_data, # 迭代初值
iter_warmup = 1000, # 每条链预处理迭代次数
iter_sampling = 2000, # 每条链总迭代次数
chains = nchains, # 马尔科夫链的数目
parallel_chains = 1, # 指定 CPU 核心数,可以给每条链分配一个
threads_per_chain = 1, # 每条链设置一个线程
show_messages = FALSE, # 不显示迭代的中间过程
refresh = 0, # 不显示采样的进度
seed = 20222022 # 设置随机数种子,不要使用 set.seed() 函数
)
# 迭代诊断
fit_poisson_stan$diagnostic_summary()$num_divergent
[1] 0 0 0 0
$num_max_treedepth
[1] 0 0 0 0
$ebfmi
[1] 1.048092 1.004069 1.061551 1.032347# A tibble: 4 × 10
variable mean median sd mad q5 q95 rhat ess_bulk
<chr> <num> <num> <num> <num> <num> <num> <num> <num>
1 alpha 0.489 0.489 0.00925 0.00926 0.474 5.04e-1 1.00 4365.
2 beta[1] 0.290 0.290 0.0148 0.0150 0.266 3.14e-1 1.00 2830.
3 beta[2] 0.215 0.215 0.0149 0.0149 0.191 2.39e-1 1.00 2941.
4 lp__ -5388. -5388. 1.22 0.993 -5390. -5.39e+3 1.00 3342.
# ℹ 1 more variable: ess_tail <num>C.4 参数后验分布
加载 bayesplot 包,可以将模型参数的后验分布图展示出来
library(ggplot2)
library(bayesplot)
mcmc_dens(fit_poisson_stan$draws(c("beta[1]", "beta[2]")),
facet_args = list(
labeller = ggplot2::label_parsed, strip.position = "top", ncol = 1
)
) +
theme_classic()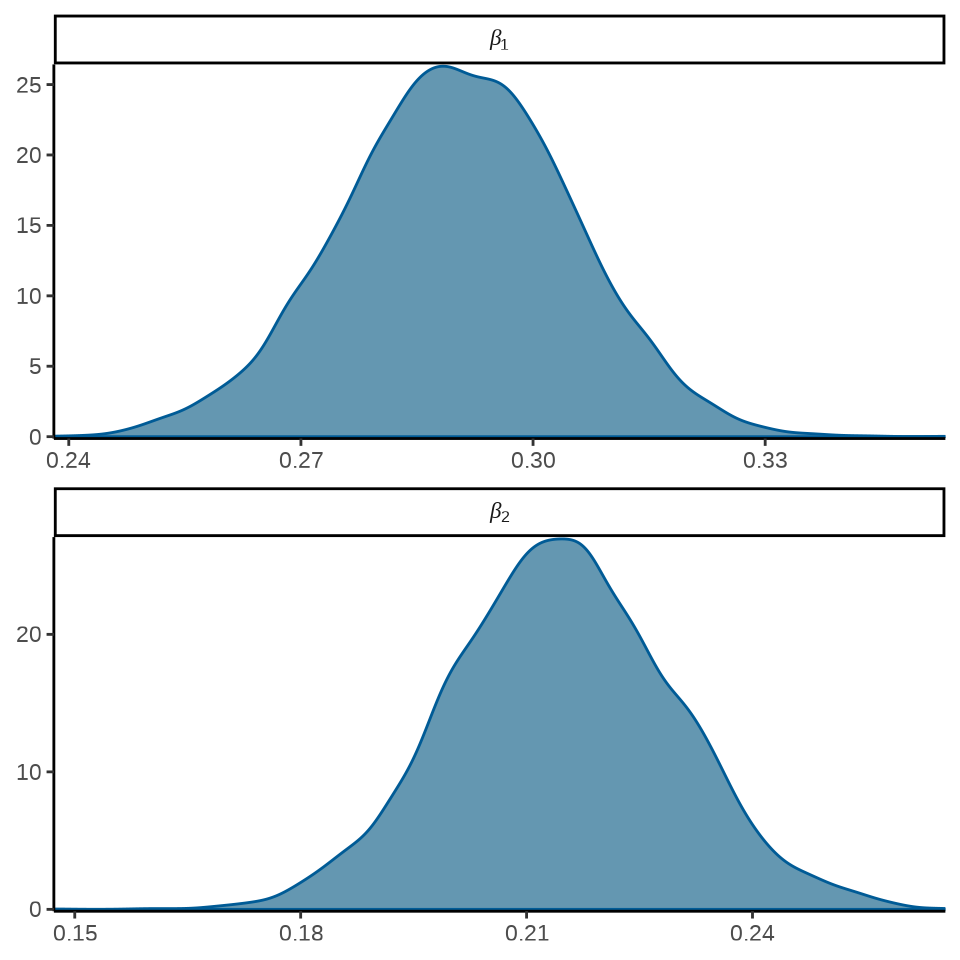
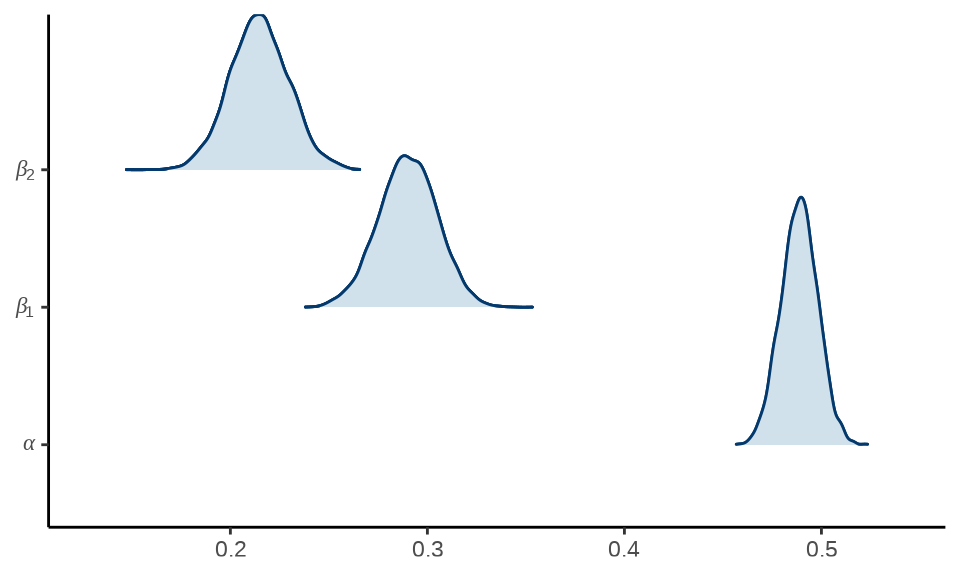
岭线图就是将各个参数的后验分布图放在一起。
C.5 模型评估指标
LOO-CV
Computed from 8000 by 2500 log-likelihood matrix
Estimate SE
elpd_loo -5386.4 37.7
p_loo 2.9 0.1
looic 10772.8 75.5
------
Monte Carlo SE of elpd_loo is 0.0.
All Pareto k estimates are good (k < 0.5).
See help('pareto-k-diagnostic') for details.C.6 brms 包
对于常见的统计模型,brms 包都内置了预编译的 Stan 程序,下面用 brms 包的函数 brm() 拟合带上述漂移项的泊松广义线性模型,参数估计结果和 Base R 函数 glm() 的几乎一致,因编译和抽样的过程比较花费时间,速度不及 Base R。
# brms
dat <- data.frame(y = y, X = X, u = u)
colnames(dat) <- c("y", "x1", "x2", "u")
fit_poisson_brm <- brms::brm(y ~ x1 + x2 + offset(log(u)),
data = dat, family = poisson(link = "log")
)
fit_poisson_brm Family: poisson
Links: mu = log
Formula: y ~ x1 + x2 + offset(log(u))
Data: dat (Number of observations: 2500)
Draws: 4 chains, each with iter = 2000; warmup = 1000; thin = 1;
total post-warmup draws = 4000
Population-Level Effects:
Estimate Est.Error l-95% CI u-95% CI Rhat Bulk_ESS Tail_ESS
Intercept 0.49 0.01 0.47 0.51 1.00 2509 2171
x1 0.29 0.01 0.26 0.32 1.00 1771 1645
x2 0.21 0.01 0.19 0.24 1.00 1727 1847
Draws were sampled using sampling(NUTS). For each parameter, Bulk_ESS
and Tail_ESS are effective sample size measures, and Rhat is the potential
scale reduction factor on split chains (at convergence, Rhat = 1).计算 LOO-CV
Computed from 4000 by 2500 log-likelihood matrix
Estimate SE
elpd_loo -5386.3 37.8
p_loo 2.9 0.1
looic 10772.6 75.5
------
Monte Carlo SE of elpd_loo is 0.0.
All Pareto k estimates are good (k < 0.5).
See help('pareto-k-diagnostic') for details.looic 指标的作用类似 AIC 指标,所以也几乎相同的。